Patterns
Deciphering Biological Networks with Patterned Heterogeneous Measurements
A modeling tool dedicated to biological network modeling (Bertrand and others 2020, <doi:10.1093/bioinformatics/btaa855>). It allows for single or joint modeling of, for instance, genes and proteins. It starts with the selection of the actors that will be the used in the reverse engineering upcoming step. An actor can be included in that selection based on its differential measurement (for instance gene expression or protein abundance) or on its time course profile. Wrappers for actors clustering functions and cluster analysis are provided. It also allows reverse engineering of biological networks taking into account the observed time course patterns of the actors. Many inference functions are provided and dedicated to get specific features for the inferred network such as sparsity, robust links, high confidence links or stable through resampling links. Some simulation and prediction tools are also available for cascade networks (Jung and others 2014, <doi:10.1093/bioinformatics/btt705>). Example of use with microarray or RNA-Seq data are provided.
README
<!-- README.md is generated from README.Rmd. Please edit that file -->
# Patterns <img src="man/figures/logo.png" align="right" width="200"/>
# A modeling tool dedicated to biological network modeling to decipher Biological Networks with Patterned Heterogeneous (e.g. multiOmics) Measurements
## Frédéric Bertrand and Myriam Maumy-Bertrand
<https://doi.org/10.32614/CRAN.package.Patterns>
<!-- badges: start -->
[](https://doi.org/10.32614/CRAN.package.Patterns)
[](https://lifecycle.r-lib.org/articles/stages.html)
[](https://www.repostatus.org/#active)
[](https://github.com/fbertran/Patterns/actions)
[](https://app.codecov.io/gh/fbertran/Patterns?branch=master)
[](https://cran.r-project.org/package=Patterns)
[](https://cran.r-project.org/package=Patterns)
[](https://github.com/fbertran/Patterns)
<!-- badges: end -->
It is designed to work with **patterned data**. Famous examples of problems related to patterned data are:
* recovering **signals** in networks after a **stimulation** (cascade network reverse engineering),
* analysing **periodic signals**.
It allows for **single** or **joint modeling** of, for instance, genes and proteins.
* It starts with the **selection of the actors** that will be the used in the reverse engineering upcoming step. An actor can be included in that selection based on its **differential effects** (for instance gene expression or protein abundance) or on its **time course profile**.
* Wrappers for **actors clustering** functions and cluster analysis are provided.
* It also allows **reverse engineering** of biological networks taking into account the observed time course patterns of the actors. Interactions between clusters of actors can be set by the user. Any number of clusters can be activated at a single time.
* Many **inference functions** are provided with the `Patterns` package and dedicated to get **specific features** for the inferred network such as **sparsity**, **robust links**, **high confidence links** or **stable through resampling links**.
+ **lasso**, from the `lars` package
+ **lasso**, from the `glmnet` package. An unweighted and a weighted version of the algorithm are available
+ **spls**, from the `spls` package
+ **elasticnet**, from the `elasticnet` package
+ **stability selection**, from the `c060` package implementation of stability selection
+ **weighted stability selection**, a new weighted version of the `c060` package implementation of stability selection that I created for the package
+ **robust**, lasso from the `lars` package with light random Gaussian noise added to the explanatory variables
+ **selectboost**, from the `selectboost` package. The selectboost algorithm looks for the more stable links against resampling that takes into account the correlated structure of the predictors
+ **weighted selectboost**, a new weighted version of the `selectboost`.
* Some **simulation** and **prediction** tools are also available for cascade networks.
* Examples of use with microarray or RNA-Seq data are provided.
The weights are viewed as a penalty factors in the penalized regression model: it is a number that multiplies the lambda value in the minimization problem to allow differential shrinkage, [Friedman et al. 2010](https://github.com/fbertran/Patterns/raw/master/add_data/glmnet.pdf), equation 1 page 3. If equal to 0, it implies no shrinkage, and that variable is always included in the model. Default is 1 for all variables. Infinity means that the variable is excluded from the model. Note that the weights are rescaled to sum to the number of variables.
A word for those that have been using our seminal work, the `Cascade` package that we created several years ago and that was a very efficient network reverse engineering tool for cascade networks
(Jung, N., Bertrand, F., Bahram, S., Vallat, L., and Maumy-Bertrand, M. (2014), <https://doi.org/10.1093/bioinformatics/btt705>, <https://cran.r-project.org/package=Cascade>, <https://github.com/fbertran/Cascade> and <https://fbertran.github.io/Cascade/>).
The `Patterns` package is more than (at least) a threeway major extension of the `Cascade` package :
* **any number of groups** can be used whereas in the `Cascade` package only 1 group for each timepoint could be created, which prevented the users to create homogeneous clusters of genes in datasets that featured more than a few dozens of genes.
* **custom** $F$ matrices shapes whereas in the `Cascade` package only 1 shape was provided:
+ interaction between groups
+ custom design of inner cells of the $F$ matrix
* the custom $F$ matrices allow to deal with **heteregeneous networks** with several kinds of actors such as mixing genes and proteins in a single network to perform **joint inference**.
* about **nine inference algorithms** are provided, whereas 1 (lasso) in `Cascade`.
Hence the `Patterns` package should be viewed more as a completely new modelling tools than as an extension of the `Cascade` package.
This website and these examples were created by F. Bertrand and M. Maumy-Bertrand.
## Installation
You can install the released version of Patterns from [CRAN](https://CRAN.R-project.org) with:
``` r
install.packages("Patterns")
```
You can install the development version of Patterns from [github](https://github.com) with:
``` r
devtools::install_github("fbertran/Patterns")
```
## Examples
### Data management
Import Cascade Data (repeated measurements on several subjects) from the CascadeData package and turn them into a omics array object. The second line makes sure the CascadeData package is installed.
``` r
library(Patterns)
```
``` r
if(!require(CascadeData)){install.packages("CascadeData")}
data(micro_US)
micro_US<-as.omics_array(micro_US[1:100,],time=c(60,90,210,390),subject=6)
str(micro_US)
#> Formal class 'omics_array' [package "Patterns"] with 7 slots
#> ..@ omicsarray: num [1:100, 1:24] 103.2 26 70.7 213.7 13.7 ...
#> .. ..- attr(*, "dimnames")=List of 2
#> .. .. ..$ : chr [1:100] "1007_s_at" "1053_at" "117_at" "121_at" ...
#> .. .. ..$ : chr [1:24] "N1_US_T60" "N1_US_T90" "N1_US_T210" "N1_US_T390" ...
#> ..@ name : chr [1:100] "1007_s_at" "1053_at" "117_at" "121_at" ...
#> ..@ gene_ID : num 0
#> ..@ group : num 0
#> ..@ start_time: num 0
#> ..@ time : num [1:4] 60 90 210 390
#> ..@ subject : num 6
```
Get a summay and plots of the data:
```r
summary(micro_US)
#> N1_US_T60 N1_US_T90 N1_US_T210 N1_US_T390
#> Min. : 12.2 Min. : 12.9 Min. : 1.5 Min. : 10.1
#> 1st Qu.: 177.7 1st Qu.: 198.7 1st Qu.: 189.0 1st Qu.: 196.7
#> Median : 513.0 Median : 499.4 Median : 608.5 Median : 541.2
#> Mean :1386.6 Mean :1357.7 Mean :1450.4 Mean :1331.2
#> 3rd Qu.:1912.3 3rd Qu.:1883.4 3rd Qu.:2050.2 3rd Qu.:1646.2
#> Max. :6348.4 Max. :6507.3 Max. :6438.5 Max. :6351.4
#> N2_US_T60 N2_US_T90 N2_US_T210 N2_US_T390
#> Min. : 16.7 Min. : 3.4 Min. : 5.5 Min. : 6.1
#> 1st Qu.: 212.4 1st Qu.: 185.7 1st Qu.: 214.7 1st Qu.: 230.1
#> Median : 584.1 Median : 501.5 Median : 596.0 Median : 601.8
#> Mean :1381.9 Mean :1345.4 Mean :1410.5 Mean :1403.7
#> 3rd Qu.:1616.2 3rd Qu.:1830.5 3rd Qu.:2005.8 3rd Qu.:1901.7
#> Max. :6149.3 Max. :6090.8 Max. :6160.6 Max. :6143.1
#> N3_US_T60 N3_US_T90 N3_US_T210 N3_US_T390
#> Min. : 1.9 Min. : 10.3 Min. : 3.3 Min. : 6.6
#> 1st Qu.: 187.4 1st Qu.: 194.6 1st Qu.: 177.8 1st Qu.: 222.6
#> Median : 611.4 Median : 576.2 Median : 552.2 Median : 593.7
#> Mean :1365.4 Mean :1381.2 Mean :1310.1 Mean :1427.1
#> 3rd Qu.:1855.2 3rd Qu.:2040.2 3rd Qu.:1784.5 3rd Qu.:2131.7
#> Max. :6636.6 Max. :6515.5 Max. :6530.4 Max. :6177.2
#> N4_US_T60 N4_US_T90 N4_US_T210 N4_US_T390
#> Min. : 20.2 Min. : 15.6 Min. : 19.8 Min. : 9.3
#> 1st Qu.: 199.3 1st Qu.: 215.4 1st Qu.: 207.0 1st Qu.: 197.8
#> Median : 610.8 Median : 614.0 Median : 544.9 Median : 590.7
#> Mean :1505.1 Mean :1526.7 Mean :1401.6 Mean :1458.8
#> 3rd Qu.:2198.1 3rd Qu.:2168.9 3rd Qu.:1831.2 3rd Qu.:1984.8
#> Max. :6986.2 Max. :7148.0 Max. :6820.0 Max. :6762.3
#> N5_US_T60 N5_US_T90 N5_US_T210 N5_US_T390
#> Min. : 3.4 Min. : 10.0 Min. : 10.7 Min. : 16.5
#> 1st Qu.: 213.2 1st Qu.: 209.8 1st Qu.: 202.0 1st Qu.: 208.2
#> Median : 609.4 Median : 561.3 Median : 555.6 Median : 570.5
#> Mean :1498.2 Mean :1424.8 Mean :1394.1 Mean :1435.3
#> 3rd Qu.:2008.7 3rd Qu.:1906.5 3rd Qu.:1923.9 3rd Qu.:1867.8
#> Max. :7268.2 Max. :6857.8 Max. :6574.0 Max. :6896.6
#> N6_US_T60 N6_US_T90 N6_US_T210 N6_US_T390
#> Min. : 13.0 Min. : 6.6 Min. : 3.8 Min. : 14.4
#> 1st Qu.: 207.5 1st Qu.: 198.6 1st Qu.: 203.9 1st Qu.: 195.8
#> Median : 516.2 Median : 530.6 Median : 578.0 Median : 580.0
#> Mean :1412.9 Mean :1388.3 Mean :1416.5 Mean :1360.8
#> 3rd Qu.:2037.4 3rd Qu.:1889.8 3rd Qu.:2030.8 3rd Qu.:1872.6
#> Max. :6898.1 Max. :6749.4 Max. :6490.0 Max. :6780.2
```
<img src="man/figures/README-plotomicsarrayclass-1.png" title="plot of chunk plotomicsarrayclass" alt="plot of chunk plotomicsarrayclass" width="100%" /><img src="man/figures/README-plotomicsarrayclass-2.png" title="plot of chunk plotomicsarrayclass" alt="plot of chunk plotomicsarrayclass" width="100%" /><img src="man/figures/README-plotomicsarrayclass-3.png" title="plot of chunk plotomicsarrayclass" alt="plot of chunk plotomicsarrayclass" width="100%" />
```r
plot(micro_US)
```
<img src="man/figures/README-plotomicsarrayclass-4.png" title="plot of chunk plotomicsarrayclass" alt="plot of chunk plotomicsarrayclass" width="100%" />
### Gene selection
There are several functions to carry out gene selection before the inference. They are detailed in the vignette of the package.
### Data simulation
Let's simulate some cascade data and then do some reverse engineering.
We first design the F matrix for $T_i=4$ times and $Ngrp=4$ groups. The `Fmat`object is an array of sizes $(T_i,T-i,Ngrp^2)=(4,4,16)$.
``` r
Ti<-4
Ngrp<-4
Fmat=array(0,dim=c(Ti,Ti,Ngrp^2))
for(i in 1:(Ti^2)){
if(((i-1) %% Ti) > (i-1) %/% Ti){
Fmat[,,i][outer(1:Ti,1:Ti,function(x,y){0<(x-y) & (x-y)<2})]<-1
}
}
```
The `Patterns` function `CascadeFinit` is an utility function to easily define such an F matrix.
``` r
Fbis=Patterns::CascadeFinit(Ti,Ngrp,low.trig=FALSE)
str(Fbis)
#> num [1:4, 1:4, 1:16] 0 0 0 0 0 0 0 0 0 0 ...
```
Check if the two matrices `Fmat` and `Fbis` are identical.
``` r
print(all(Fmat==Fbis))
#> [1] TRUE
```
End of F matrix definition.
``` r
Fmat[,,3]<-Fmat[,,3]*0.2
Fmat[3,1,3]<-1
Fmat[4,2,3]<-1
Fmat[,,4]<-Fmat[,,3]*0.3
Fmat[4,1,4]<-1
Fmat[,,8]<-Fmat[,,3]
```
We set the seed to make the results reproducible and draw a scale free random network.
``` r
set.seed(1)
Net<-Patterns::network_random(
nb=100,
time_label=rep(1:4,each=25),
exp=1,
init=1,
regul=round(rexp(100,1))+1,
min_expr=0.1,
max_expr=2,
casc.level=0.4
)
Net@F<-Fmat
str(Net)
#> Formal class 'omics_network' [package "Patterns"] with 6 slots
#> ..@ omics_network: num [1:100, 1:100] 0 0 0 0 0 0 0 0 0 0 ...
#> ..@ name : chr [1:100] "gene 1" "gene 2" "gene 3" "gene 4" ...
#> ..@ F : num [1:4, 1:4, 1:16] 0 0 0 0 0 0 0 0 0 0 ...
#> ..@ convF : num [1, 1] 0
#> ..@ convO : num 0
#> ..@ time_pt : int [1:4] 1 2 3 4
```
Plot the simulated network.
``` r
Patterns::plot(Net, choice="network")
```
If a gene clustering is known, it can be used as a coloring scheme.
``` r
plot(Net, choice="network", gr=rep(1:4,each=25))
```
Plot the F matrix, for low dimensional F matrices.
``` r
plot(Net, choice="F")
```
<div class="figure">
<img src="man/figures/README-plotF-1.png" alt="plot of chunk plotF" width="100%" />
<p class="caption">plot of chunk plotF</p>
</div>
Plot the F matrix using the `pixmap` package, for high dimensional F matrices.
``` r
plot(Net, choice="Fpixmap")
```
<div class="figure">
<img src="man/figures/README-plotFpixmap-1.png" alt="plot of chunk plotFpixmap" width="100%" />
<p class="caption">plot of chunk plotFpixmap</p>
</div>
We simulate gene expression according to the network that was previously drawn
``` r
set.seed(1)
M <- Patterns::gene_expr_simulation(
network=Net,
time_label=rep(1:4,each=25),
subject=5,
peak_level=200,
act_time_group=1:4)
#> Error: unable to find an inherited method for function 'gene_expr_simulation' for signature 'omics_network = "missing"'
str(M)
#> num [1:60, 1:6] -1.495 0.368 0.517 -0.484 0.675 ...
```
Get a summay and plots of the simulated data:
```r
summary(M)
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'object' lors de la s�lection d'une m�thode pour la fonction 'summary' : objet 'M' introuvable
```
```r
plot(M)
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'x' lors de la s�lection d'une m�thode pour la fonction 'plot' : objet 'M' introuvable
```
### Network inferrence
We infer the new network using subjectwise leave one out cross-validation (default setting): all measurements from the same subject are removed from the dataset). The inference is carried out with a general Fshape.
```r
Net_inf_P <- Patterns::inference(M, cv.subjects=TRUE)
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P' not found
```
Heatmap of the inferred coefficients of the Omega matrix
``` r
stats::heatmap(Net_inf_P@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P' not found
```
Default values fot the $F$ matrices. The `Finit` matrix (starting values for the algorithm). In our case, the `Finit`object is an array of sizes $(T_i,T-i,Ngrp^2)=(4,4,16)$.
``` r
Ti<-4;
ngrp<-4
nF<-ngrp^2
Finit<-array(0,c(Ti,Ti,nF))
for(ii in 1:nF){
if((ii%%(ngrp+1))==1){
Finit[,,ii]<-0
} else {
Finit[,,ii]<-cbind(rbind(rep(0,Ti-1),diag(1,Ti-1)),rep(0,Ti))+rbind(cbind(rep(0,Ti-1),diag(1,Ti-1)),rep(0,Ti))
}
}
```
The `Fshape` matrix (default shape for `F` matrix the algorithm). Any interaction between groups and times are permitted except the retro-actions (a group on itself, or an action at the same time for an actor on another one).
``` r
Fshape<-array("0",c(Ti,Ti,nF))
for(ii in 1:nF){
if((ii%%(ngrp+1))==1){
Fshape[,,ii]<-"0"
} else {
lchars <- paste("a",1:(2*Ti-1),sep="")
tempFshape<-matrix("0",Ti,Ti)
for(bb in (-Ti+1):(Ti-1)){
tempFshape<-replaceUp(tempFshape,matrix(lchars[bb+Ti],Ti,Ti),-bb)
}
tempFshape <- replaceBand(tempFshape,matrix("0",Ti,Ti),0)
Fshape[,,ii]<-tempFshape
}
}
```
Any other form can be used. A "0" coefficient is missing from the model. It allows testing the best structure of an "F" matrix and even performing some significance tests of hypothses on the structure of the $F$ matrix.
The `IndicFshape` function allows to design custom F matrix for cascade networks with equally spaced measurements by specifying the zero and non zero $F_{ij}$ cells of the $F$ matrix. It is useful for models featuring several clusters of actors that are activated at the time. Let's define the following indicatrix matrix (action of all groups on each other, which is not a possible real modeling setting and is only used as an example):
``` r
TestIndic=matrix(!((1:(Ti^2))%%(ngrp+1)==1),byrow=TRUE,ngrp,ngrp)
TestIndic
#> [,1] [,2] [,3] [,4]
#> [1,] FALSE TRUE TRUE TRUE
#> [2,] TRUE FALSE TRUE TRUE
#> [3,] TRUE TRUE FALSE TRUE
#> [4,] TRUE TRUE TRUE FALSE
```
For that choice, we get those init and shape $F$ matrices.
``` r
IndicFinit(Ti,ngrp,TestIndic)
#> , , 1
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 0 0 0 0
#> [3,] 0 0 0 0
#> [4,] 0 0 0 0
#>
#> , , 2
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 3
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 4
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 5
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 6
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 0 0 0 0
#> [3,] 0 0 0 0
#> [4,] 0 0 0 0
#>
#> , , 7
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 8
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 9
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 10
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 11
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 0 0 0 0
#> [3,] 0 0 0 0
#> [4,] 0 0 0 0
#>
#> , , 12
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 13
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 14
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 15
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 1 0 0 0
#> [3,] 1 1 0 0
#> [4,] 1 1 1 0
#>
#> , , 16
#>
#> [,1] [,2] [,3] [,4]
#> [1,] 0 0 0 0
#> [2,] 0 0 0 0
#> [3,] 0 0 0 0
#> [4,] 0 0 0 0
IndicFshape(Ti,ngrp,TestIndic)
#> , , 1
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "0" "0" "0" "0"
#> [3,] "0" "0" "0" "0"
#> [4,] "0" "0" "0" "0"
#>
#> , , 2
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 3
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 4
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 5
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 6
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "0" "0" "0" "0"
#> [3,] "0" "0" "0" "0"
#> [4,] "0" "0" "0" "0"
#>
#> , , 7
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 8
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 9
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 10
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 11
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "0" "0" "0" "0"
#> [3,] "0" "0" "0" "0"
#> [4,] "0" "0" "0" "0"
#>
#> , , 12
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 13
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 14
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 15
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "a1" "0" "0" "0"
#> [3,] "a2" "a1" "0" "0"
#> [4,] "a3" "a2" "a1" "0"
#>
#> , , 16
#>
#> [,1] [,2] [,3] [,4]
#> [1,] "0" "0" "0" "0"
#> [2,] "0" "0" "0" "0"
#> [3,] "0" "0" "0" "0"
#> [4,] "0" "0" "0" "0"
```
Those $F$ matrices are lower diagonal ones to enforce that an observed value at a given time can only be predicted by a value that was observed in the past only (i.e. neither at the same moment or in the future).
The `plotF` is convenient to display F matrices. Here are the the displays of the three $F$ matrices we have just introduced.
``` r
plotF(Fshape,choice="Fshape")
```
<div class="figure">
<img src="man/figures/README-plotfshape1-1.png" alt="plot of chunk plotfshape1" width="100%" />
<p class="caption">plot of chunk plotfshape1</p>
</div>
``` r
plotF(CascadeFshape(4,4),choice="Fshape")
```
<div class="figure">
<img src="man/figures/README-plotfshape2-1.png" alt="plot of chunk plotfshape2" width="100%" />
<p class="caption">plot of chunk plotfshape2</p>
</div>
``` r
plotF(IndicFshape(Ti,ngrp,TestIndic),choice="Fshape")
```
<div class="figure">
<img src="man/figures/README-plotfshape3-1.png" alt="plot of chunk plotfshape3" width="100%" />
<p class="caption">plot of chunk plotfshape3</p>
</div>
We now fit the model with an $F$ matrix that is designed for cascade networks.
Specific Fshape
```r
Net_inf_P_S <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4))
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_S, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_S' not found
```
Heatmap of the coefficients of the Omega matrix of the network. They reflect the use of a special $F$ matrix. It is an example of an F matrix specifically designed to deal with cascade networks.
``` r
stats::heatmap(Net_inf_P_S@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_S' not found
```
There are many fitting functions provided with the `Patterns` package in order to search for **specific features** for the inferred network such as **sparsity**, **robust links**, **high confidence links** or **stable through resampling links**. :
* **LASSO**, from the `lars` package
* **LASSO2**, from the `glmnet` package. An unweighted and a weighted version of the algorithm are available
* **SPLS**, from the `spls` package
* **ELASTICNET**, from the `elasticnet` package
* **stability.c060**, from the `c060` package implementation of stability selection
* **stability.c060.weighted**, a new weighted version of the `c060` package implementation of stability selection
* **robust**, lasso from the `lars` package with light random Gaussian noise added to the explanatory variables
* **selectboost.weighted**, a new weighted version of the `selectboost` package implementation of the selectboost algorithm to look for the more stable links against resampling that takes into account the correlated structure of the predictors. If no weights are provided, equal weigths are for all the variables (=non weighted case).
```r
Net_inf_P_Lasso2 <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4), fitfun="LASSO2")
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_Lasso2, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_Lasso2' not found
```
Heatmap of the coefficients of the Omega matrix of the network
``` r
stats::heatmap(Net_inf_P_Lasso2@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_Lasso2' not found
```
We create a weighting vector to perform weighted lasso inference.
``` r
Weights_Net=slot(Net,"network")
#> Error in slot(Net, "network"): no slot of name "network" for this object of class "omics_network"
Weights_Net[Net@network!=0]=.1
#> Error: object 'Weights_Net' not found
Weights_Net[Net@network==0]=1000
#> Error: object 'Weights_Net' not found
```
```r
Net_inf_P_Lasso2_Weighted <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4), fitfun="LASSO2", priors=Weights_Net)
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_Lasso2_Weighted, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_Lasso2_Weighted' not found
```
Heatmap of the coefficients of the Omega matrix of the network
``` r
stats::heatmap(Net_inf_P_Lasso2_Weighted@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_Lasso2_Weighted' not found
```
```r
Net_inf_P_SPLS <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4), fitfun="SPLS")
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_SPLS, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_SPLS' not found
```
Heatmap of the coefficients of the Omega matrix of the network
``` r
stats::heatmap(Net_inf_P_SPLS@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_SPLS' not found
```
```r
Net_inf_P_ELASTICNET <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4), fitfun="ELASTICNET")
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_ELASTICNET, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_ELASTICNET' not found
```
Heatmap of the coefficients of the Omega matrix of the network
``` r
stats::heatmap(Net_inf_P_ELASTICNET@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_ELASTICNET' not found
```
```r
Net_inf_P_stability <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4), fitfun="stability.c060")
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_stability, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_stability' not found
```
Heatmap of the coefficients of the Omega matrix of the network
``` r
stats::heatmap(Net_inf_P_stability@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_stability' not found
```
```r
Net_inf_P_StabWeight <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4), fitfun="stability.c060.weighted", priors=Weights_Net)
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_StabWeight, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_StabWeight' not found
```
Heatmap of the coefficients of the Omega matrix of the network
``` r
stats::heatmap(Net_inf_P_StabWeight@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_StabWeight' not found
```
```r
Net_inf_P_Robust <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4), fitfun="robust")
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_Robust, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_Robust' not found
```
Heatmap of the coefficients of the Omega matrix of the network
``` r
stats::heatmap(Net_inf_P_Robust@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_Robust' not found
```
```r
Weights_Net_1 <- Weights_Net
#> Error in eval(expr, envir, enclos): objet 'Weights_Net' introuvable
Weights_Net_1[,] <- 1
#> Error in Weights_Net_1[, ] <- 1: objet 'Weights_Net_1' introuvable
Net_inf_P_SelectBoost <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4), fitfun="selectboost.weighted",priors=Weights_Net_1)
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
```
#>
#> Attaching package: 'Patterns'
#> The following object is masked from 'package:testthat':
#>
#> compare
#> The following object is masked from 'package:igraph':
#>
#> compare
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_SelectBoost, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_SelectBoost' not found
```
Heatmap of the coefficients of the Omega matrix of the network
``` r
stats::heatmap(Net_inf_P_SelectBoost@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_SelectBoost' not found
```
```r
Net_inf_P_SelectBoostWeighted <- Patterns::inference(M, Finit=CascadeFinit(4,4), Fshape=CascadeFshape(4,4), fitfun="selectboost.weighted",priors=Weights_Net)
#> Error in h(simpleError(msg, call)): erreur d'�valuation de l'argument 'M' lors de la s�lection d'une m�thode pour la fonction 'inference' : objet 'M' introuvable
```
```
#>
#> Attaching package: 'Patterns'
#> The following object is masked from 'package:testthat':
#>
#> compare
#> The following object is masked from 'package:igraph':
#>
#> compare
```
Plot of the inferred F matrix
``` r
plot(Net_inf_P_SelectBoostWeighted, choice="F")
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'plot': object 'Net_inf_P_SelectBoostWeighted' not found
```
Heatmap of the coefficients of the Omega matrix of the network
``` r
stats::heatmap(Net_inf_P_SelectBoostWeighted@network, Rowv = NA, Colv = NA, scale="none", revC=TRUE)
#> Error: object 'Net_inf_P_SelectBoostWeighted' not found
```
### Post inference network analysis
Such an analysis is only required if the model was not fitted using the stability selection or the selectboost algorithm.
Create an animation of the network with increasing cutoffs with an animated .gif format or a html webpage in the working directory.
``` r
data(network)
sequence<-seq(0,0.2,length.out=20)
evolution(network,sequence,type.ani = "gif", outdir=getwd())
evolution(network,sequence,type.ani = "html", outdir=getwd())
```
```
#> Error in setwd(outdir): cannot change working directory
#> Error in setwd(outdir): cannot change working directory
```

[Evolution as .html.](https://fbertran.github.io/Patterns/reference/evolution/index.html)
Evolution of some properties of a reverse-engineered network with increasing cut-off values.
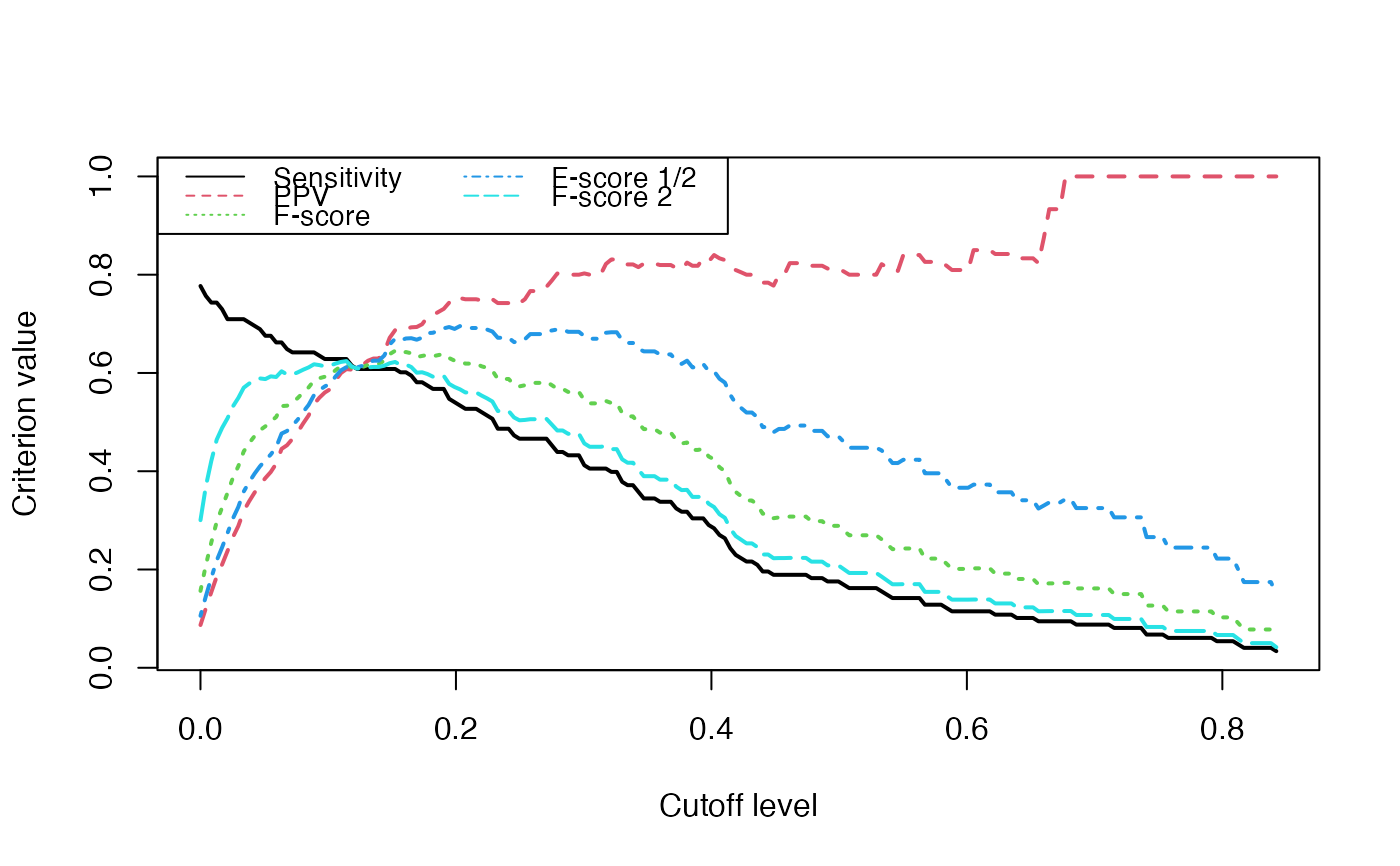
We switch to data that were derived from the inferrence of a real biological network and try to detect the optimal cutoff value: the best cutoff value for a network to fit a scale free network. The `cutoff` was validated only single group cascade networks (number of actors groups = number of timepoints) and for genes dataset. Instead of the `cutoff` function, manual curation or the stability selection or the selectboost algorithm should be used.
```r
data("networkCascade")
set.seed(1)
cutoff(networkCascade)
#> Error in (function (classes, fdef, mtable) : impossible de trouver une méthode héritée pour la fonction 'cutoff' pour la signature '"network"'
```
Analyze the network with a cutoff set to the previouly found 0.133 optimal value.
```r
analyze_network(networkCascade,nv=0.133)
#> Error in (function (classes, fdef, mtable) : impossible de trouver une méthode héritée pour la fonction 'analyze_network' pour la signature '"network"'
```
``` r
data(Selection)
plot(networkCascade,nv=0.133, gr=Selection@group)
```
<div class="figure">
<img src="man/figures/README-plotnet-1.png" alt="plot of chunk plotnet" width="100%" />
<p class="caption">plot of chunk plotnet</p>
</div>
### Perform gene selection
Import data.
``` r
library(Patterns)
library(CascadeData)
data(micro_S)
micro_S<-as.omics_array(micro_S,time=c(60,90,210,390),subject=6,gene_ID=rownames(micro_S))
data(micro_US)
micro_US<-as.omics_array(micro_US,time=c(60,90,210,390),subject=6,gene_ID=rownames(micro_US))
```
Select early genes (t1 or t2):
```r
Selection1<-geneSelection(x=micro_S,y=micro_US,20,wanted.patterns=rbind(c(0,1,0,0),c(1,0,0,0),c(1,1,0,0)))
```
Section genes with first significant differential expression at t1:
```r
Selection2<-geneSelection(x=micro_S,y=micro_US,20,peak=1)
```
Section genes with first significant differential expression at t2:
```r
Selection3<-geneSelection(x=micro_S,y=micro_US,20,peak=2)
```
Select later genes (t3 or t4)
```r
Selection4<-geneSelection(x=micro_S,y=micro_US,50,
wanted.patterns=rbind(c(0,0,1,0),c(0,0,0,1),c(1,1,0,0)))
```
Merge those selections:
``` r
Selection<-unionOmics(Selection1,Selection2)
Selection<-unionOmics(Selection,Selection3)
Selection<-unionOmics(Selection,Selection4)
head(Selection)
#> $omicsarray
#> US60 US90 US210
#> 210226_at 0.82417544 0.9166931 0.7310784
#> 233516_s_at -0.27395188 -2.3695246 0.6511830
#> 202081_at 0.60477249 0.6599672 -0.1884742
#> 236719_at -2.07284086 -0.3123747 0.1792494
#> 236019_at -0.08175065 -0.3699708 -0.4315901
#> 1563563_at -1.44513486 1.6869516 -0.4297297
#>
#> $name
#> [1] "210226_at" "233516_s_at" "202081_at" "236719_at" "236019_at" "1563563_at"
#>
#> $gene_ID
#> [1] "210226_at" "233516_s_at" "202081_at" "236719_at" "236019_at" "1563563_at"
#>
#> $group
#> [1] 1 2 1 1 1 1
#>
#> $start_time
#> [1] 1 2 1 1 1 1
#>
#> $time
#> [1] 60 90 210 390
#>
#> $subject
#> [1] 6
```
Summarize the final selection:
``` r
summary(Selection)
#> US60 US90 US210 US390
#> Min. :-2.76841 Min. :-2.369525 Min. :-1.6147 Min. :-2.60480
#> 1st Qu.:-0.18028 1st Qu.:-0.181425 1st Qu.: 0.1985 1st Qu.:-0.03884
#> Median : 0.05675 Median :-0.001924 Median : 0.9886 Median : 0.31766
#> Mean : 0.05764 Mean : 0.278275 Mean : 0.9611 Mean : 0.26428
#> 3rd Qu.: 0.22438 3rd Qu.: 0.664063 3rd Qu.: 1.6918 3rd Qu.: 0.57117
#> Max. : 2.86440 Max. : 4.284675 Max. : 3.6727 Max. : 2.54704
#> US60 US90 US210 US390
#> Min. :-2.7932 Min. :-2.49245 Min. :-1.21606 Min. :-1.74407
#> 1st Qu.:-0.5547 1st Qu.:-0.01944 1st Qu.: 0.07967 1st Qu.:-0.26548
#> Median :-0.3089 Median : 0.14977 Median : 0.72019 Median : 0.03616
#> Mean :-0.2917 Mean : 0.33720 Mean : 0.73063 Mean : 0.06753
#> 3rd Qu.:-0.1725 3rd Qu.: 0.48744 3rd Qu.: 1.26164 3rd Qu.: 0.32496
#> Max. : 2.0267 Max. : 3.37588 Max. : 3.87950 Max. : 2.83321
#> US60 US90 US210 US390
#> Min. :-2.94444 Min. :-0.9721 Min. :-1.9349 Min. :-3.8418
#> 1st Qu.:-0.23136 1st Qu.:-0.1027 1st Qu.: 0.3254 1st Qu.:-0.1592
#> Median :-0.04761 Median : 0.2548 Median : 1.2512 Median : 0.1538
#> Mean : 0.22116 Mean : 0.6479 Mean : 1.0485 Mean : 0.1219
#> 3rd Qu.: 0.33157 3rd Qu.: 1.0737 3rd Qu.: 1.8513 3rd Qu.: 0.6268
#> Max. : 3.31723 Max. : 4.3604 Max. : 4.4860 Max. : 1.9886
#> US60 US90 US210 US390
#> Min. :-2.85438 Min. :-0.90355 Min. :-0.83324 Min. :-0.96834
#> 1st Qu.:-0.06031 1st Qu.:-0.08464 1st Qu.: 0.07605 1st Qu.: 0.01569
#> Median : 0.03601 Median : 0.17135 Median : 0.52176 Median : 0.17370
#> Mean : 0.14593 Mean : 0.41929 Mean : 0.62446 Mean : 0.23854
#> 3rd Qu.: 0.24568 3rd Qu.: 0.75564 3rd Qu.: 1.07821 3rd Qu.: 0.45189
#> Max. : 1.82903 Max. : 3.60640 Max. : 2.27744 Max. : 1.90880
#> US60 US90 US210 US390
#> Min. :-1.38002 Min. :-2.94444 Min. :-1.0271 Min. :-1.3636
#> 1st Qu.:-0.19910 1st Qu.:-0.01758 1st Qu.: 0.1459 1st Qu.:-0.1386
#> Median :-0.07962 Median : 0.16080 Median : 0.7430 Median : 0.1492
#> Mean : 0.12972 Mean : 0.37123 Mean : 0.7972 Mean : 0.1271
#> 3rd Qu.: 0.26113 3rd Qu.: 0.61933 3rd Qu.: 1.3922 3rd Qu.: 0.4825
#> Max. : 2.31074 Max. : 3.24454 Max. : 3.6213 Max. : 1.5979
#> US60 US90 US210 US390
#> Min. :-1.79176 Min. :-3.20791 Min. :-1.4716 Min. :-1.95883
#> 1st Qu.:-0.09822 1st Qu.:-0.03963 1st Qu.: 0.1292 1st Qu.:-0.04786
#> Median : 0.03378 Median : 0.28261 Median : 0.8392 Median : 0.22472
#> Mean : 0.27978 Mean : 0.52529 Mean : 0.7903 Mean : 0.21171
#> 3rd Qu.: 0.33548 3rd Qu.: 1.03256 3rd Qu.: 1.4416 3rd Qu.: 0.42511
#> Max. : 3.16035 Max. : 3.19975 Max. : 2.8027 Max. : 2.14903
```
<div class="figure">
<img src="man/figures/README-omicsselection6-1.png" alt="plot of chunk omicsselection6" width="100%" />
<p class="caption">plot of chunk omicsselection6</p>
</div><div class="figure">
<img src="man/figures/README-omicsselection6-2.png" alt="plot of chunk omicsselection6" width="100%" />
<p class="caption">plot of chunk omicsselection6</p>
</div><div class="figure">
<img src="man/figures/README-omicsselection6-3.png" alt="plot of chunk omicsselection6" width="100%" />
<p class="caption">plot of chunk omicsselection6</p>
</div>
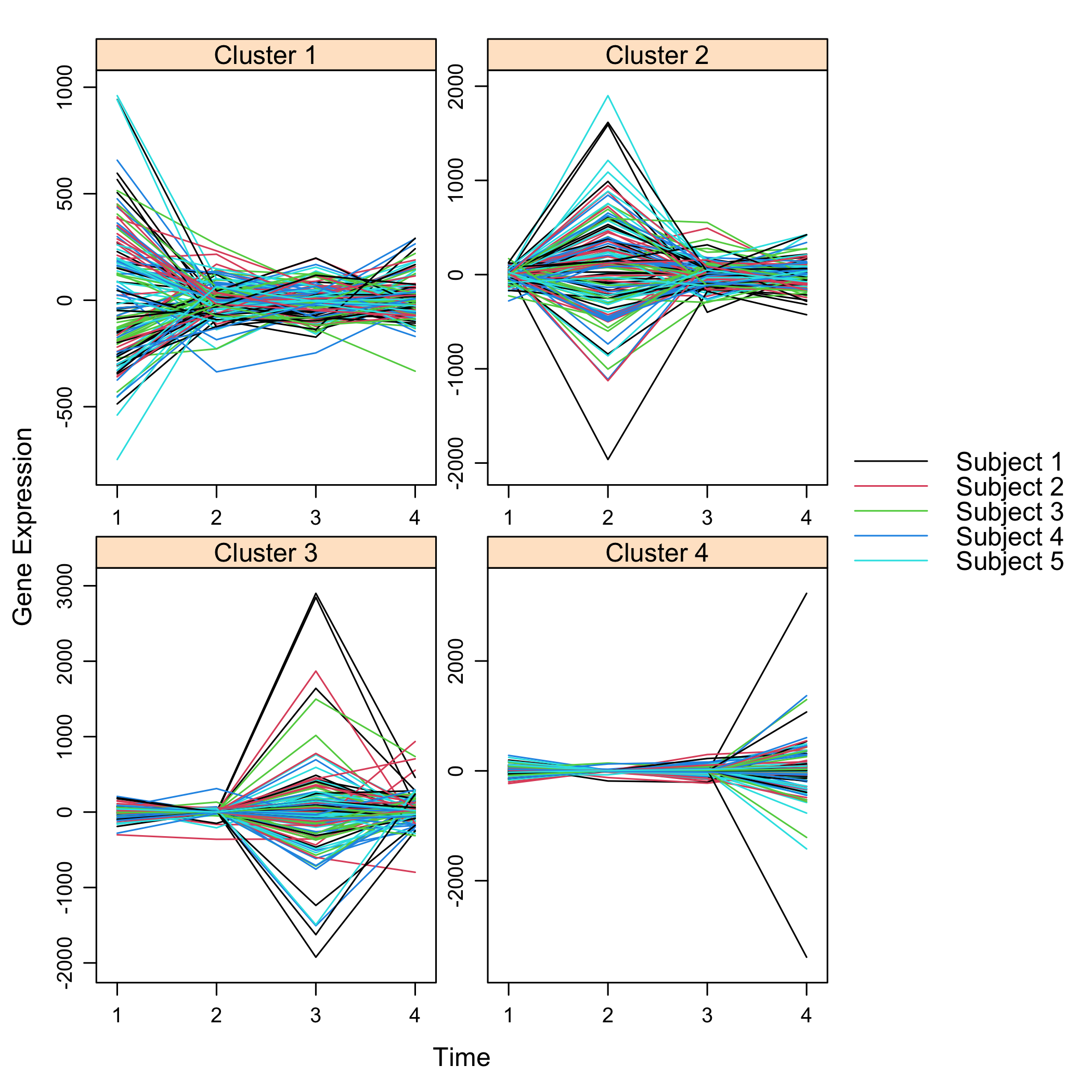
Plot the final selection:
``` r
plot(Selection)
```
<div class="figure">
<img src="man/figures/README-omicsselection7-1.png" alt="plot of chunk omicsselection7" width="100%" />
<p class="caption">plot of chunk omicsselection7</p>
</div><div class="figure">
<img src="man/figures/README-omicsselection7-2.png" alt="plot of chunk omicsselection7" width="100%" />
<p class="caption">plot of chunk omicsselection7</p>
</div><div class="figure">
<img src="man/figures/README-omicsselection7-3.png" alt="plot of chunk omicsselection7" width="100%" />
<p class="caption">plot of chunk omicsselection7</p>
</div><div class="figure">
<img src="man/figures/README-omicsselection7-4.png" alt="plot of chunk omicsselection7" width="100%" />
<p class="caption">plot of chunk omicsselection7</p>
</div><div class="figure">
<img src="man/figures/README-omicsselection7-5.png" alt="plot of chunk omicsselection7" width="100%" />
<p class="caption">plot of chunk omicsselection7</p>
</div><div class="figure">
<img src="man/figures/README-omicsselection7-6.png" alt="plot of chunk omicsselection7" width="100%" />
<p class="caption">plot of chunk omicsselection7</p>
</div><div class="figure">
<img src="man/figures/README-omicsselection7-7.png" alt="plot of chunk omicsselection7" width="100%" />
<p class="caption">plot of chunk omicsselection7</p>
</div><div class="figure">
<img src="man/figures/README-omicsselection7-8.png" alt="plot of chunk omicsselection7" width="100%" />
<p class="caption">plot of chunk omicsselection7</p>
</div>
This process could be improved by retrieve a real gene_ID using the `bitr` function of the `ClusterProfiler` package or by performing independent filtering using `jetset` package to only keep at most only probeset (the best one, if there is one good enough) per gene_ID.
### Examples of outputs
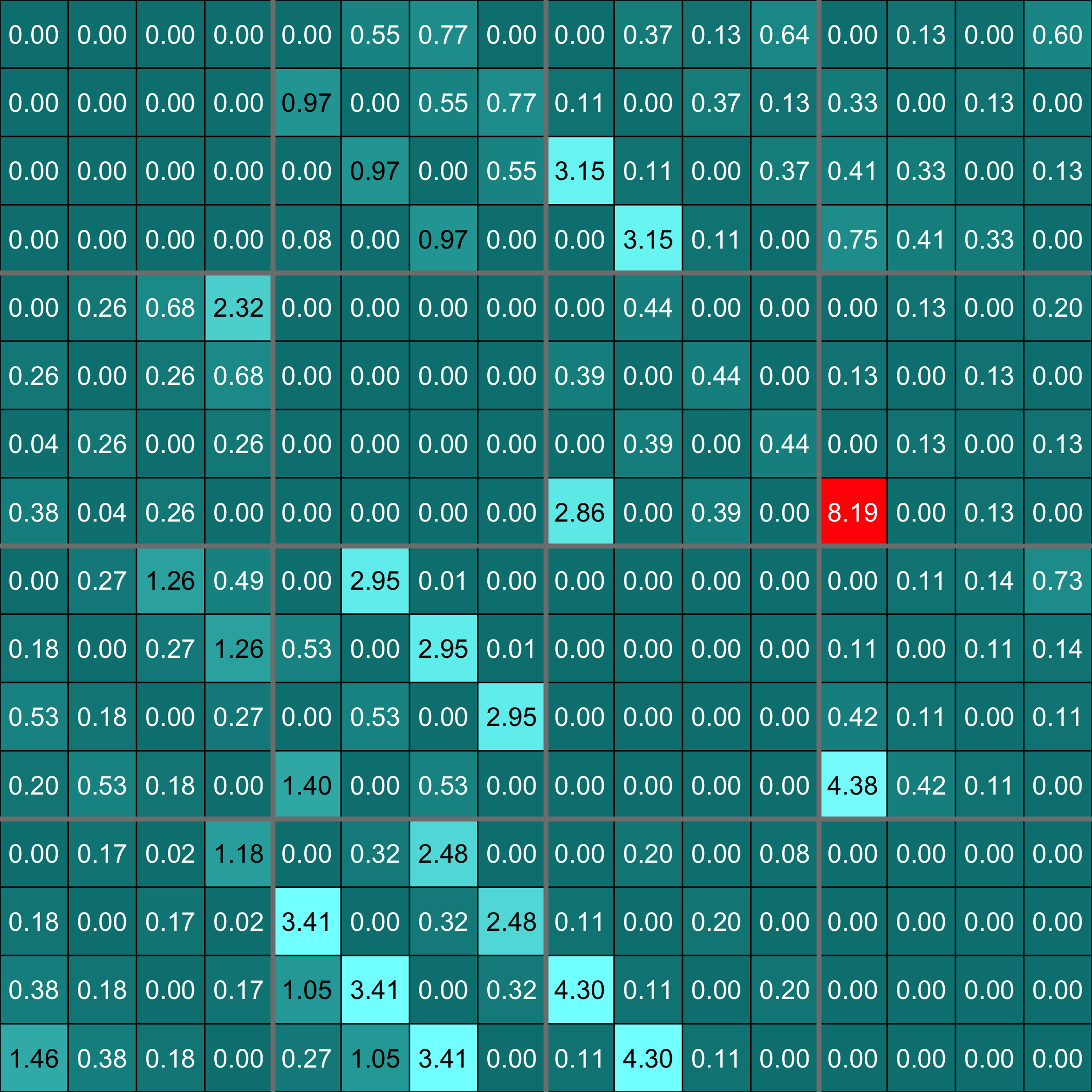
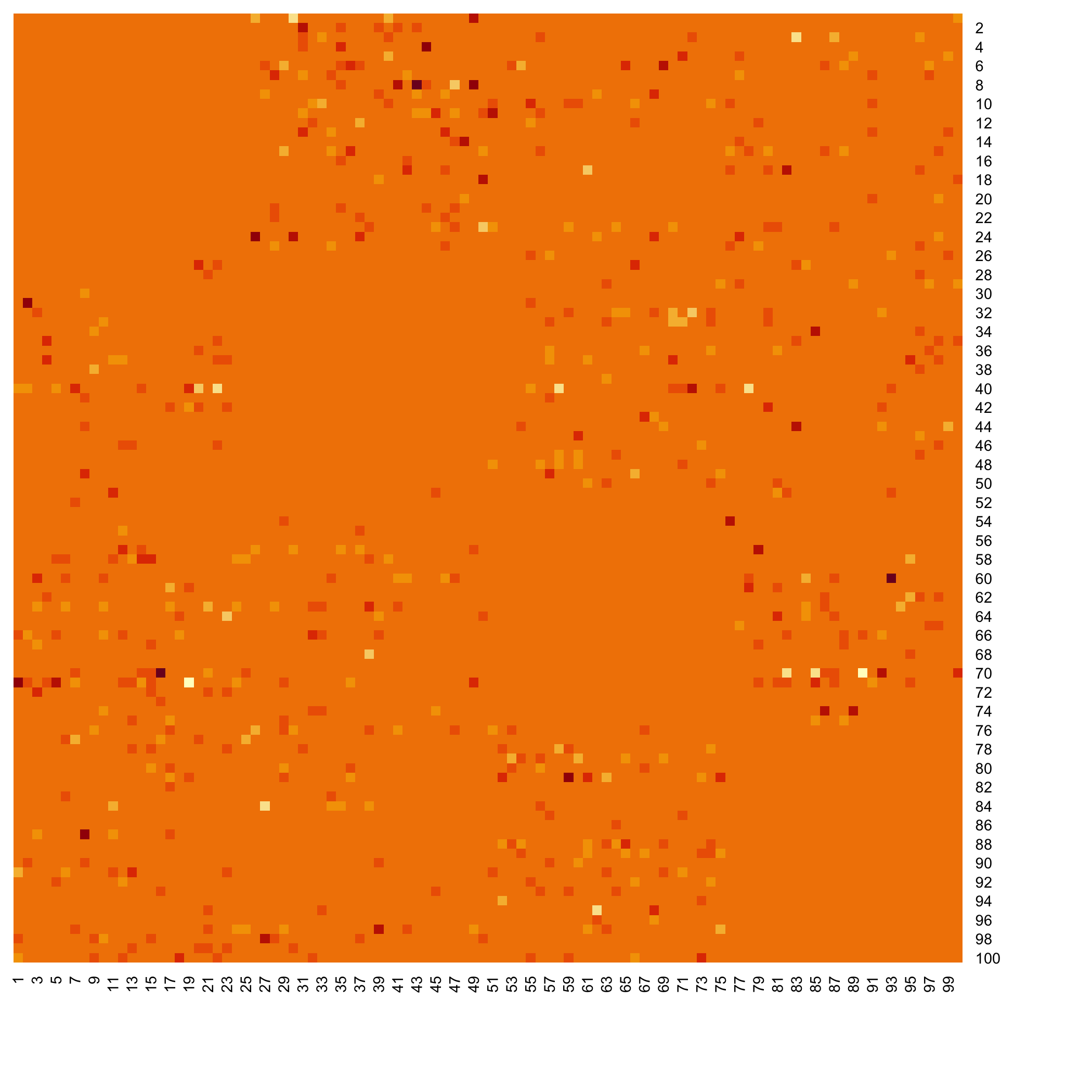



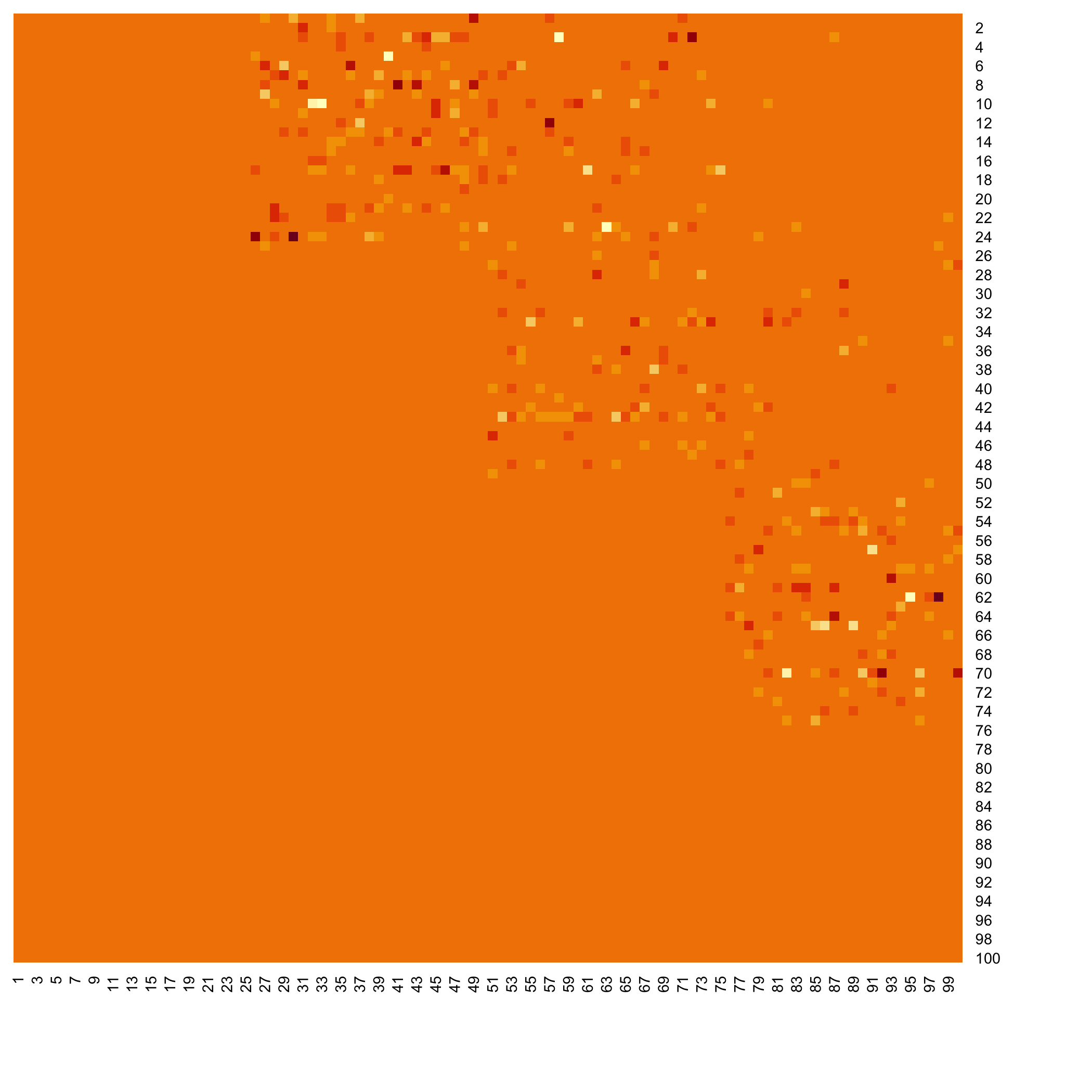
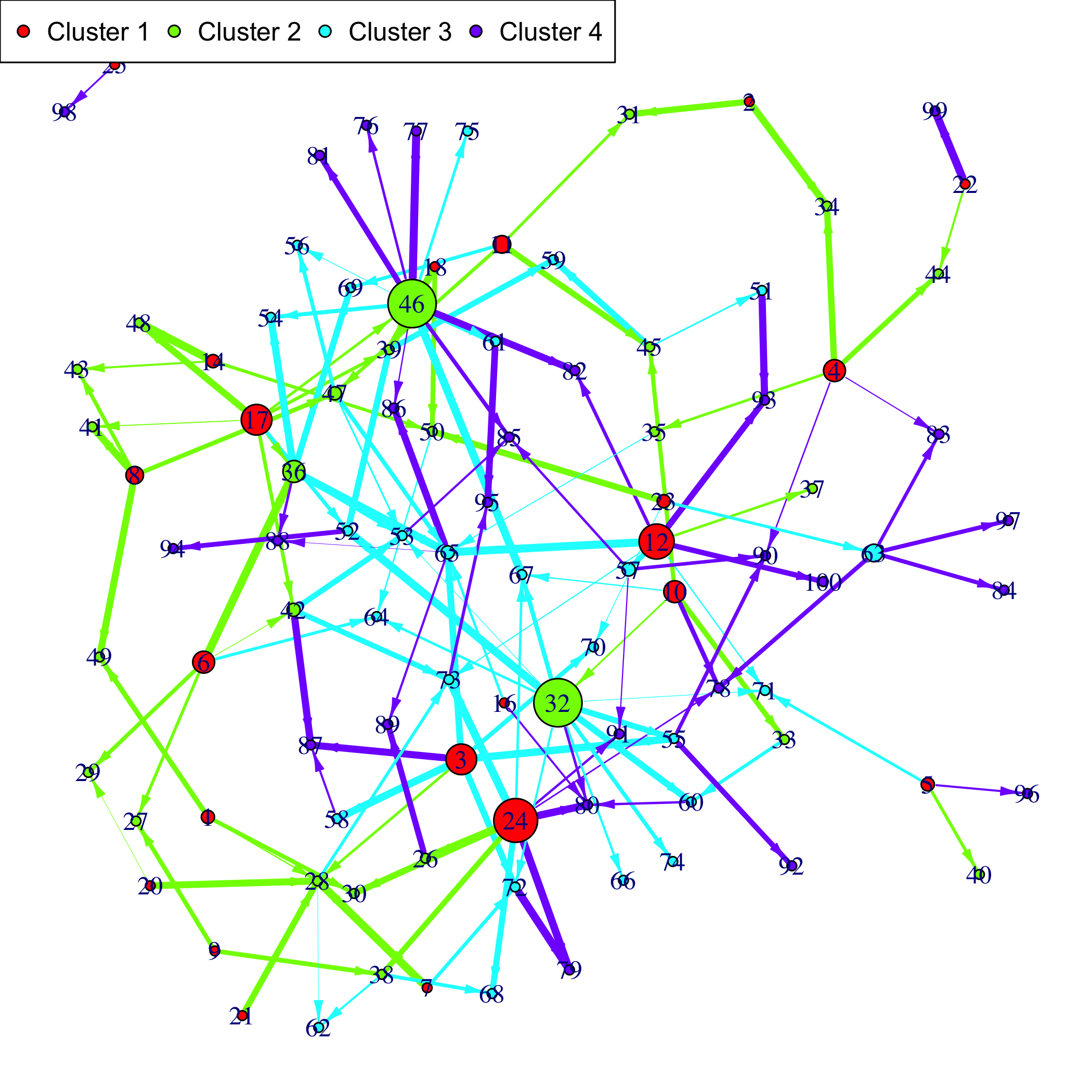

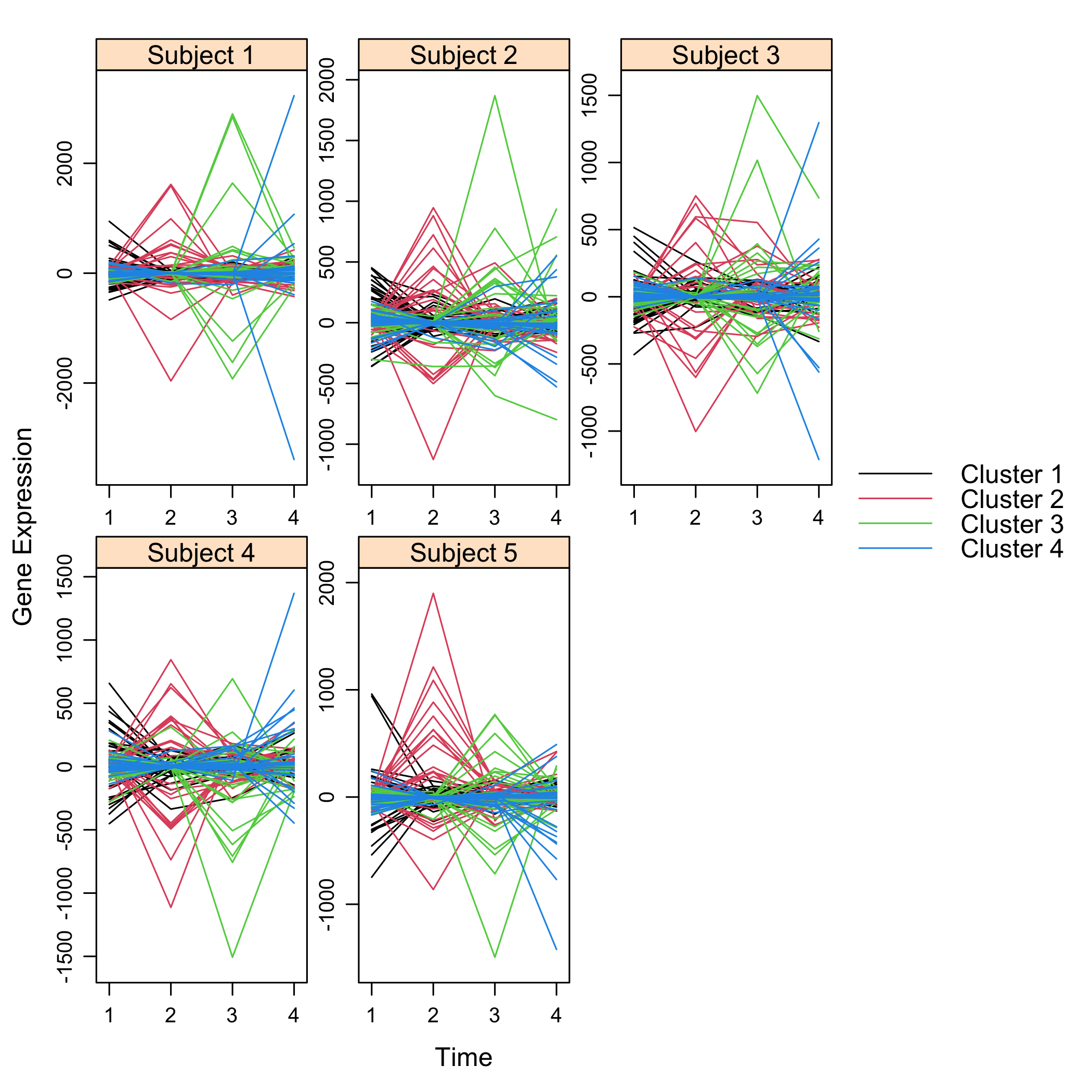
Versions across snapshots
| Version | Repository | File | Size |
|---|---|---|---|
1.7 |
rolling linux/jammy R-4.5 | Patterns_1.7.tar.gz |
1006.0 KiB |
1.7 |
rolling linux/noble R-4.5 | Patterns_1.7.tar.gz |
1006.0 KiB |
1.7 |
rolling source/ R- | Patterns_1.7.tar.gz |
1006.0 KiB |
1.7 |
latest linux/jammy R-4.5 | Patterns_1.7.tar.gz |
1006.0 KiB |
1.7 |
latest linux/noble R-4.5 | Patterns_1.7.tar.gz |
1006.0 KiB |
1.7 |
latest source/ R- | Patterns_1.7.tar.gz |
1006.0 KiB |
1.7 |
2026-04-26 source/ R- | Patterns_1.7.tar.gz |
1006.0 KiB |
1.7 |
2026-04-23 source/ R- | Patterns_1.7.tar.gz |
1006.0 KiB |
1.5 |
2025-04-20 source/ R- | Patterns_1.5.tar.gz |
1.3 MiB |